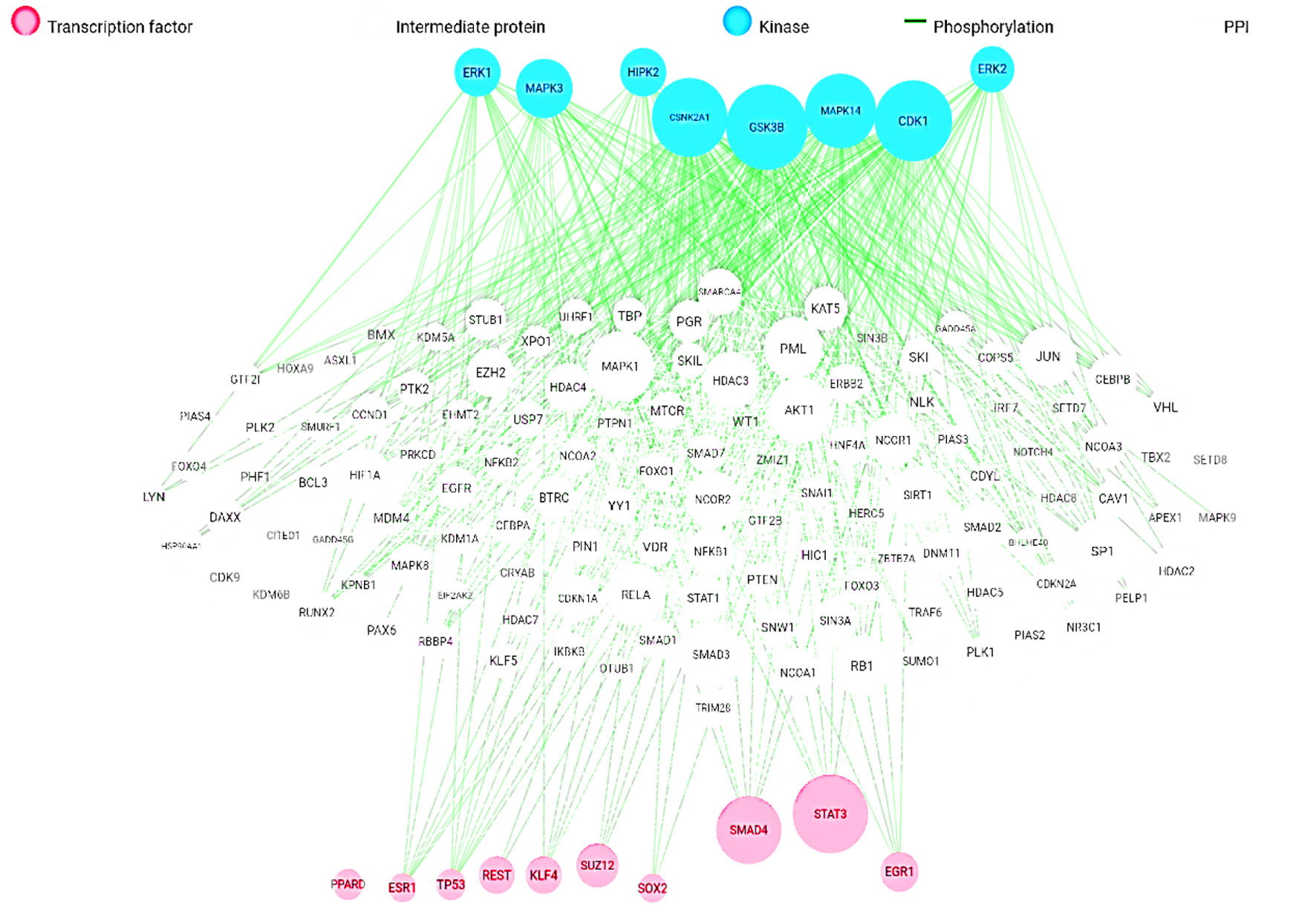
Figure 1. Overall target genes network.
| Journal of Food Bioactives, ISSN 2637-8752 print, 2637-8779 online |
| Journal website www.isnff-jfb.com |
Original Research
Volume 23, September 2023, pages 58-67
In Silico Evaluation of Nutri-Pharmacological Potentials of Phytochemicals in Sorghum (Sorghum bicolor) Grains
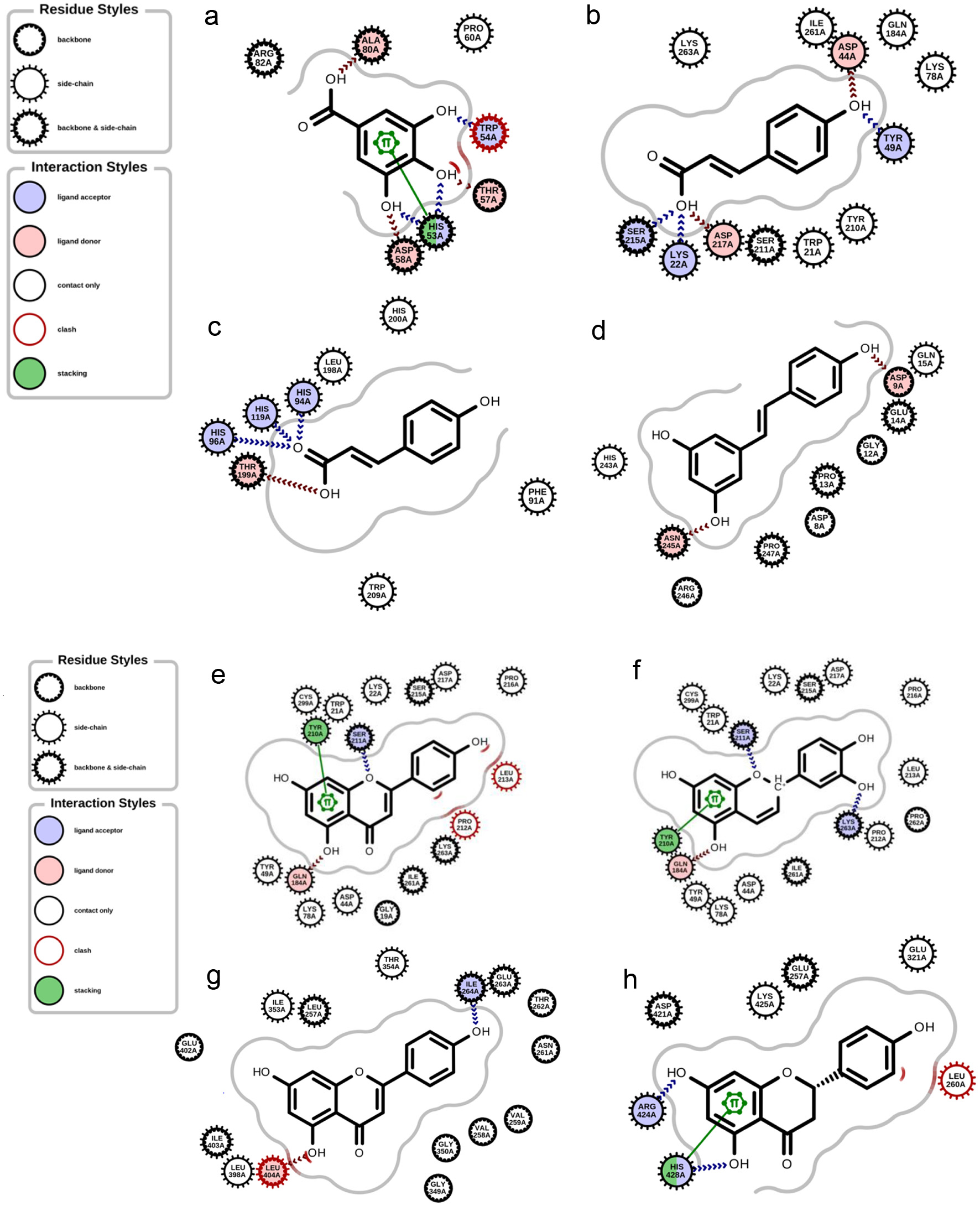
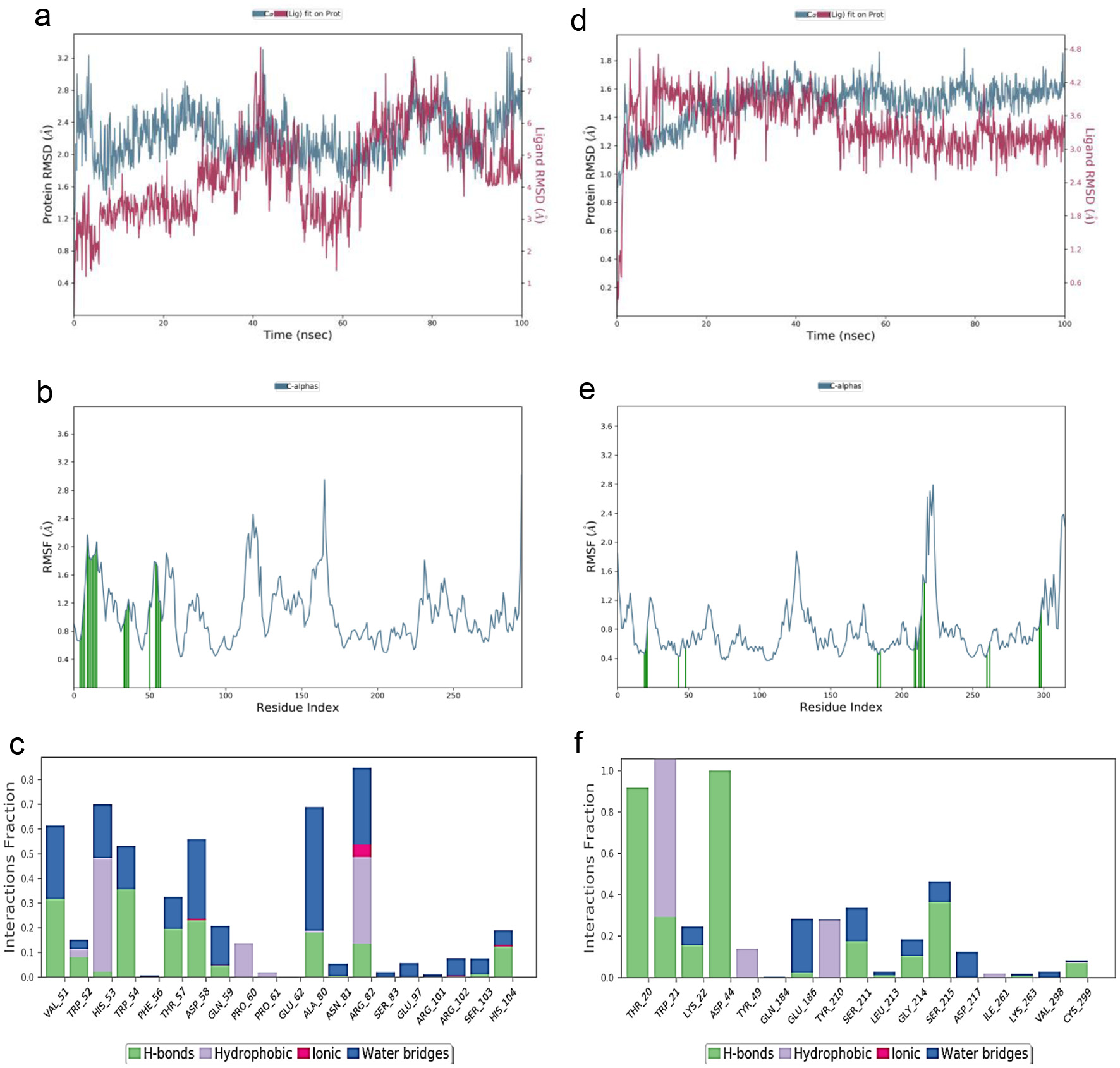
Figures

Tables
| SN | Ligand | PUBCHEM ID | MW | MR | TPSA (Å2) | LOGP | ESOL LOGS | ESOL CLASS | GIA | BBB | P-GP | CYPS INHIBITOR | LOG Kp (CM/S) | BS | SA |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Physicochemical properties: Molecular weight (MW), Molar Refractivity (MR), Total polar surface area (TPSA). Pharmacokinetics: Gastrointestinal absorption (GIA), Blood-brain barrier (BBB), P-glycoprotein (P-gp) substrate, Inhibition of Cytochrome P450 (CYPs) type CYP1A2, CYP2C19, CYP2C9, CYP2D6, and CYP3A4, Skin permeation (Log Kp). Water Solubility: ESOL Log S, ESOL Class. Lipophilicity: Consensus Log P. Medicinal Chemistry: Synthetic accessibility (SA). Druglikeness: Bioavailability Score (BS). | |||||||||||||||
| 1 | Protocatechuic acid | 528594 | 154.12 | 37.45 | 77.76 | 0.65 | −1.85 | Very Soluble | High | No | No | CYP3A4 | −6.42 | 0.56 | 1.07 |
| 2 | p-hydroxybenzoic acid | 135 | 138.12 | 35.42 | 57.53 | 1.05 | −2.07 | Soluble | High | Yes | No | None | −6.02 | 0.85 | 1.00 |
| 3 | Vanillic acid | 8468 | 168.15 | 41.92 | 66.76 | 1.08 | −2.02 | Soluble | High | No | No | None | −6.31 | 0.85 | 1.42 |
| 4 | p-coumaric acid. | 1549106 | 164.16 | 45.13 | 57.53 | 1.26 | −2.02 | Soluble | High | Yes | No | None | −6.26 | 0.85 | 1.61 |
| 5 | Ferulic acid | 445858 | 194.18 | 51.63 | 66.76 | 1.36 | −2.11 | Soluble | High | Yes | No | None | −6.41 | 0.85 | 1.93 |
| 6 | Gallic acid | 24721416 | 170.12 | 39.47 | 97.99 | 0.21 | −164 | Soluble | High | No | No | None | −6.84 | 0.56 | 1.22 |
| 7 | Caffeic acid | 154911 | 180.16 | 47.16 | 77.76 | 0.93 | −1.89 | Very Soluble | High | No | No | None | −6.58 | 0.56 | 1.81 |
| 8 | Cinnamic acid | 53729554 | 148.16 | 43.11 | 37.30 | 1.79 | −2.37 | Soluble | High | Yes | No | None | −5.69 | 0.85 | 1.67 |
| 9 | Salicyclic acid. | 46782907 | 138.12 | 35.42 | 57.53 | 1.24 | −2.50 | Soluble | High | Yes | No | None | −5.54 | 0.85 | 1.00 |
| 10 | Syringic acid | 10742 | 75.99 | 0.99 | −1.84 | Very Soluble | High | No | No | None | −6.77 | 0.56 | 1.70 | ||
| 11 | Sinapic acid | 637775 | 224.21 | 58.12 | 75.99 | 1.31 | −2.16 | Soluble | High | No | No | None | −6.00 | 0.56 | 2.17 |
| 12 | Apigeninidin. | 159360 | 271.24 | 74.15 | 94.06 | 0.85 | −2.94 | Soluble | High | No | Yes | CYP1A2 | −6.95 | 0.55 | 2.92 |
| 13 | Luteolinidin. | 441701 | 270.24 | 73.99 | 90.90 | 2.11 | −3.94 | Soluble | High | No | No | CYP1A2, CYP2D6 | −5.80 | 0.55 | 2.96 |
| 14 | Apigenin. | 528044 | 272.25 | 71.57 | 86.99 | 1.84 | −3.49 | Soluble | High | No | Yes | CYP1A2, CYP2D6, CYP3A4 | −6.17 | 0.55 | 3.01 |
| 15 | Luteolin | 5280445 | 288.25 | 73.59 | 1.45 | 1.45 | −3.26 | Soluble | High | No | Yes | CYP1A2, CYP2D6, CYP3A4 | −6.62 | 0.55 | 3.11 |
| 16 | Trans-resveratrol | 71684637 | 390.38 | 100 | 139.84 | 0.62 | −2.90 | Soluble | High | No | Yes | None | −7.95 | 0.55 | 2.02 |
| 17 | Naringenin | 932 | 272.25 | 71.57 | 86.99 | 1.84 | −3.49 | Soluble | High | No | Yes | CYP1A2, CYP3A4 | −6.17 | 0.55 | 3.01 |
| SN | Ligands | % Probability of Predicted Targets | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | ||
| A. Carbon anhydrase I/II/VI/VII/XII (P00915/P00918/P23280/P43166/O43570). B. Carbon anhydrase III/IV/VA/VB/IX/XI/XIII/XIV (P07451/P22748/P35218/Q16790/Q8N1Q1/Q9ULX7), C. Aldose reductase (P15121), D. Estrogen receptor (Q92731), E. Macrophage migration inhibitory factor (P14174), F. Alpha (1–3)-fucosyltransferase7 (Q11130), G. Arachidonate 5-lioxygenase (P09917), H. Matrix metaloprotenase9 (P14780), I. Matrix metaloprotenase1 (P03956), J. Matrix metaloprotenase2 (P08253). K. Protein-tyrosine phosphatase 1B (P18031). L. Hydroxycarboxylic acid receptor (Q8TDS4). M. Lymphocyte differentiation antigen (P28907). N. NADPH Oxidase 4 (Q9NPH5). O. Cyclin-dependent kinase 5 (Q00535). P. Xanthine dehydrogenase (P47989). | |||||||||||||||||
| 1 | Protocatechuic acid | 98 | 98 | ||||||||||||||
| 2 | p-hydroxy benzoic acid | 100 | 100 | ||||||||||||||
| 3 | Vanillic acid | 30 | 25 | 30 | 25 | ||||||||||||
| 4 | p-coumaric acid | 100 | 100 | 100 | 100 | ||||||||||||
| 5 | Ferulic acid | 98 | 98 | ||||||||||||||
| 6 | Gallic acid | 100 | 100 | 100 | |||||||||||||
| 7 | Caffeic acid | 80 | 80 | 80 | 80 | 80 | 80 | 80 | |||||||||
| 8 | Trans resveratrol | 100 | |||||||||||||||
| 9 | Salicylic acid | 100 | 25 | ||||||||||||||
| 10 | Syringic acid | 100 | 100 | ||||||||||||||
| 11 | Sinapic acid | 25 | 25 | ||||||||||||||
| 12 | Luteolinidin | 100 | 100 | ||||||||||||||
| 13 | Apigenin | 100 | 100 | 100 | 100 | 100 | |||||||||||
| 14 | Luteolin | 80 | |||||||||||||||
| 15 | Naringenin | 95 | 95 | 95 | |||||||||||||
| 16 | Salicyclic acid | 100 | 40 | ||||||||||||||
| 17 | Cinnamic acid | 90 | |||||||||||||||
| SN | Protein | Binding Affinity Score(kcal.mol−1) | ||||||
|---|---|---|---|---|---|---|---|---|
| p-Coumaric acid | Gallic acid | Trans-Resveratrol | Apigenin | Luteolinidin | Naringenin | |||
| Docking parameter: Arachidonate 5-lioxygenase [Center: −4.57, 2.552, −4.052; Size: 126, 106, 126; Spacing 0.867]. Aldose reductase [Center: 1.346, 2.385, 0.255; Size:126, 126, 126; Spacing: 0.400]. Carbon anhydrase I [Center: 36.650, 14.797, −13.116; Size:126, 126, 126; Spacing: 0.375]. Alpha (1–3) fucosynltransferase 7 [Center: 0.064, 2.743, 7.051; Size: 76, 70, 126; Spacing: 1.000]. Carbon anhydrase IV [Center: −3.911, 2.015, −4.599; Size: 126, 126, 68; Spacing: 0.844]. Carbon anhydrase XII [Center: −6.047, 0.271, −6.751; Size:126, 120, 126; Spacing: 0.536]. | ||||||||
| 1 | Arachidonate 5-lioxygenase | −5.620 | ||||||
| 2 | Aldose reductase | −7.759 | −10.430 | −10.500 | ||||
| 3 | Carbon anhydrase I | −5.554 | −5.982 | |||||
| 4 | Alpha (1–3) fucosynltransferase 7 | −5.512 | ||||||
| 5 | Carbon anhydrase IV | −5.163 | ||||||
| 6 | Carbon anhydrase XII | −6.328 | ||||||
| 7 | Estrogen receptor | −7.513 | −6.648 | −6.778 | ||||
| 8 | NADPH oxidase 4 | −8.287 | ||||||
| 9 | Cyclin dependent kinase 5 | −8.366 | ||||||
| 10 | Xanthine dehydrogenase | −9.196 | ||||||
| Complex | Simulation Time (ns) | MMGBSA ΔGbind (kcal.mol−1) | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Coulomb | Covalent | Hbond | Lipo | Packing | Solv_GB | vdW | Gbind (Total) | ||
| Legend: Covalent: Covalent binding energy. Coulomb: Coulomb energy. Lipo: Lipophilic energy. Hbond: Hydrogen bonding energy. Packing: Pi-pi packing correction. Solv GB: Generalized Born electrostatic solvation energy. vdW: Van der Waals energy. Total: Total energy (Prime energy). | |||||||||
| p-Coumaric acid and aldose reductase | 0 | −13.980 | 1.333 | −1.308 | −22.041 | −0.002 | 12.445 | −33.076 | −56.631 |
| 100 | −16.204 | 5.018 | −1.244 | −21.909 | −0.002 | 12.689 | −31.892 | −53.546 | |
| Gallic acid and alpha-(1,3)-fucosyltransferase 7 | 0 | −19.226 | 2.005 | −0.554 | −8.601 | −2.273 | 18.761 | −19.743 | −29.632 |
| 100 | −14.336 | 2.594 | −0.845 | −7.264 | −0.767 | 10.884 | −18.929 | −28.663 | |